POTENTIAL APPLICATIONS OF MODERN BIOLOGICAL TECHNIQUES IN BREEDING FRUIT TREES
|
|
|
- Weronika Szczepaniak
- 7 lat temu
- Przeglądów:
Transkrypt
1 Journal of Fruit and Ornamental Plant Research Vol. 14 (Suppl. 1), 2006 POTENTIAL APPLICATIONS OF MODERN BIOLOGICAL TECHNIQUES IN BREEDING FRUIT TREES U r i La v i 1 *, S l av a Gu r e vitz 1, Gi a r a Ben A ri 2, D a vi d Saada 1, K h alib K as hkush 1, T al i P az 1, T e T wi to 1, Y u v al Cohen 1, J ossi Hillel 2 an d Gi o ra Simch en 2 1 Institute of Horticulture, ARO-Volcani Center. P.O.B. 6 Bet-Dagan, 50250, ISRAEL 2 The Hebrew University Faculty of Agriculture Rehovot and Institute of Life Sciences, Jerusalem, ISRAEL *Corresponding author: ulavi@volcani.agri.gov.il (Received July 15, 2005/Accepted October 30, 2005) A B S T R A C T The current methodologies of fruit tree breeding are based on segregating population and/or induced mutation, and the selection of the desired phenotype. Looking at limitations such as long juvenile period, large tree size, many not applicable technologies (backcrossing and hybrid seeds), classical breeding is expensive and very time-consuming, which makes the development of better and more efficient techniques an urgent priority. Modern biological techniques not only can be used to breed new fruit tree cultivars, but also can be applied to many other aspects of the fruit growing industry. Modern biological techniques are not yet widely used in fruit plant breeding. One reason is that the genetics of fruit trees is still not well understood. Another reason is that reliable and efficient transformation protocols have not yet been developed for most fruit tree species. This manuscript focuses on a few techniques which are currently available for application in fruit trees. These techniques can be used to identify important genes which can be used to generate transgenic plants. They can also be used in Marker Assisted Selection. These new genetic techniques include DNA chips and bioinformatics. Key words: AFLP markers, SNP analysis, gene introgression, off type plants, biodiversity
2 U. Lavi et al. INTRODUCT ION There is no question that classical breeding techniques are currently the only ones used to produce new fruit tree cultivars. Most currently available fruit tree cultivars were developed either by randomly identifying selected seedlings or by using classic breeding techniques in traditional breeding programs. But these techniques have many limitations, which makes the development of better and more efficient techniques an urgent priority. Modern biological techniques not only can be used to breed new fruit tree cultivars, but also can be applied to many other aspects of the fruit growing industry. Many different strategies support classical breeding and cultivation. Some transgenic plants have already been planted on a commercial scale and have made a significant impact on both current and future agriculture. Nevertheless, the real revolution has barely started, not so much because of public mistrust of genetically modified crops, but rather because the techniques currently available are still limited. Transgenic maize, rice and cotton cultivars have been quit successful. The cultivars have been modified to be resistant to herbicides, insects or viruses. Golden rice is a modified cultivar with enhanced nutritional value. However, only one transformed fruit tree cultivar is commercially available: a papaya cultivar which is resistant to PRV. One reason is that the genetics of fruit trees is still not well understood. Another reason is that reliable and efficient transformation protocols have not yet been developed for most fruit tree species. The second way of modern breeding is development of molecular markers. DNA markers are DNA fragments that differ among the varieties of a given species. Marker profiles generated by various molecular techniques can be used to identify and distinguish cultivars and rootstocks. They can be used to enhance the efficiency of classic breeding programs. Identification by DNA markers can also be used to protect the legal rights of breeders. A few examples of how our group has used DNA markers to improve fruit breeding and cultivation are presented below. 1. Mango malformation Mango malformation is a devastating disease, which can cause enormous economical loss almost everywhere mangoes are grown. It is caused by the fungus Fusarium mangiferae. Inflorescences of infected trees are malformed and unable to bear fruit. The epidemiology of the disease is not yet well understood. No effective treatments or reliable diagnostic tools are currently available. Because the disease affects only the inflorescences, infection is extremely difficult to detect when the tree is not in bloom. A diagnostic tool is needed to not only to keep orchards free of the pathogen, but also to better understand the epidemiology of the disease. We have developed a simple PCR based method. Using AFLP, we identified and 14 J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006: 13-19
3 Potential applications of modern biological tech niques in breeding screened pathogen-specific DNA sequences to ensure that they did not occur either in other Fusarium species or in the mango genome itself. We then sequenced this fragments and designed primers for PCR. The procedure works well in the laboratory, and is being improved in terms of both specificity and sensitivity. Trials in commercial orchards are already underway. 2. Decreasing the number of generations needed for backcrossing after gene introgression Backcrossing is a common breeding technique to introduce a specific gene from a wild variety into a commercial cultivar. Six to eight backcrosses are generally needed to dilute out the undesired wild variety genes. In species which take several years to mature, backcrossing can take decades. This drastically limits the usefulness of backcrossing in breeding fruit trees. We have proposed using DNA marker techniques to reduce the number of backcross generations needed (Hillel et al., 1990). Even after the first backcross, the percentage of the progeny genome which is derived from the recurrent parent s genome ranges from 50 to 100%. DNA marker techniques can identify those individuals which carry the target gene and have the highest proportion of the recurrent parent genome. Depending on the number of chromosomes and the level of selection, the number of backcrosses can be reduced dramatically to one to three generations. The initial investment of screening a large number of progeny can pay back handsomely in saved time. 3. Mango Amplified Fragment Length Polymorphism (AFLP) is a DNA fingerprinting technique based on the selective PCR amplification of restriction fragments from a total digest of genomic DNA. The technique can be used with any species and does not require any prior information on DNA sequences. Fifty to one hundred restriction fragments are typically amplified and visualized on denaturing polyacrylamide gels (Vos et al., 1995). We have used AFLP to generate a preliminary genetic map of the mango and to assess the genetic relationships between various mango cultivars and rootstocks (Kashkush et al., 2001). 4. Date palms Modern propagation of date palms is based on tissue culture. This technique is essential for the rapid propagation of a large number of trees and is the basis for the modern date palm industry. However, propagation by tissue culture frequently produces off-types, some of which are detected only after the plants reach maturity. This can cause major economic losses for the growers. J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006:
4 U. Lavi et al. In order to detect and prevent off-types, propagation protocols need to be modified based on a thorough understanding of the mechanism by which offtypes arise. We have been using AFLP in order to broaden our knowledge of this phenomenon. We compared the AFLP band patterns of normal and off-type trees of the cultivars Barhee and Medjoul. We identified many sequence polymorphisms among the Medjoul trees which had been produced by tissue culture. Nevertheless, we did not find any specific AFLP band associated with a specific off-type. Therefore, we used methylation-sensitive and methylation-resistant restriction enzymes to compare methylation patterns in normal and off-type trees. Preliminary results suggest that each off-type has a characteristic methylation pattern. This had already been found to be the case in oil palms as well. We are currently using cdna-aflp to compare differences in gene expression between normal and off-type trees. We are also studying several genes in which changes giving rise to off-types seem to occur with particular frequency. Among the changes being studied are changes in DNA sequence, methylation, and gene expression (Gurevitz et al., 2004). 5. Yeast a. Single Nucleotide Polymorphism (SNP) SNP s are the most common polymorphisms in the human genome and most probably in other genomes as well. Other types of sequence variation, such as chromosomal aberrations and variation in repeat number occur far less often. They do not seem to be randomly distributed throughout the genome. SNP s are under intensive investigation as potential tools to assess genetic variation and identify genes. These point mutations account for most of the genetic variation among individuals, and have thus become important tools in medical genetics and evolutionary biology. Almost 10 million human SNP s have been described and are being used in medical research, population studies and forensic science. SNP s occur in high numbers in the genomes of all organisms. High throughput techniques have been developed to ensure rapid and accurate SNP genotyping. b. Using SNP s to study biodiversity and hunt for genes in yeast Saccharomyces cerevlsiae is an excellent model organism for obtaining knowledge about genetic processes which sometimes can be applied to other organisms, including fruit trees. We have used SNP s to identify yeast genes responsible for differences in sporulation efficiency. We studied two strains of S. cerevisiae: SK1, which has high sporulation efficiency, and S288c, which has low sporulation 16 J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006: 13-19
5 Potential applications of modern biological tech niques in breeding efficiency. A total of 145 genes consisting of 81,480 base pairs are known to be associated with sporulation in S. cerevisiae. By sequencing these genes in the SK1 genome, 554 SNP s were found which distinguished SK1 from S288c. Sporulation efficiency was determined in SK1, S288c, the F1 hybrid, and 326 diploid segregants generated from meiosis in the F1 hybrid. Sporulation efficiency was 90% in SK1, and 15% in S288c. In the hybrid, sporulation efficiency was closer to SK1 than to S228c. In the segregants, sporulation efficiency ranged from 1% to 97.5%, with an average of 69%. Two pools of DNA were prepared: one from 21 segregants with the highest sporulation efficiencies, and the other from 21 segregants with the lowest sporulation efficiencies. Allele frequencies were calculated for each SNP in both groups by visualizing the two alleles inherited from SK1 and S288c. The only genes in which significant differences in allele frequency were found were YNL100W and RAS2, which are located 3 Kb apart on chromosome 14. This "candidate region" was found to be 175 Kb in length. Genome-wide screening was performed by hybridizing the DNA of the two strains and the two segregant DNA pools to the S98 Affymetrix yeast chip. We identified about 4000 "probes" containing SNP s distinguishing SK1 from S288c. Regions which contained alleles which differed between the two segregant pools were also detected by hybridization. Three regions containing several probes were identified. On the basis of the candidate gene search and the genome-wide screen, we decided to focus our attention on the candidate region on chromosome 14. Based on earlier research, we chose fourteen candidate genes which might play a role in sporulation or meiosis. These genes were tested to see if they contained single mutations which might account for the difference in sporulation between SK1 and S288c. Reciprocal homozygosity was also investigated. Four such genes were found. We have verified our results using several different techniques: 1. Analysis of double and quadruple heterozygote deletions. 2. Comparative sequence analysis. A specific deletion of one A in a poly-a stretch in genes RAS2 and SWS2 was found to distinguish between SK1 and S288c. The deletion was located between the TATA box and the first codon. These findings suggest that the deletion could be the mutation responsible for the differences in sporulation efficiency between SK1 and S288c. 3. Analysis of expression patterns in the two alleles of RAS2 and SWS2 by Real Time PCR. Expression of RAS2 seems to be the same in the two alleles. However, expression of SWS2 is significantly different, especially at time zero, when there is a 2.5 fold difference. 4. Allele replacement. Allele replacement was performed using both the SK1 alleles of RAS2 and SWS2. The alleles were introduced into S288c. Background and sporulation efficiencies were measured. Sporulation efficiency was 0.7% in the S288c transformed with the decreasing allele of J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006:
6 U. Lavi et al. the RAS2 gene, 17.1% in the wild type, and 50.1% in the S288c transformed with SWS2. 6. Concluding remarks Our results suggest that genes controlling traits of interest in fruit trees might be identified by techniques such as searching for candidate genes and sequencing DNA pools. In order to definitively assign a function to a given gene, transformation with the specific allele has to be accompanied by a change in phenotype. Transformation protocols have already been developed for some fruit trees and are being developed for others. Some of the methods used by human geneticists may also be of use in assigning functions to genes in fruit trees species. REFERENCES Hillel J., Schaap T., Haberfeld A., Jeffreys A.J., Plotsky Y., Cahane R.A., Lavi U DNA Fingerprints applied to gene introgression in breeding programs. GENETICS 124: Kashkush K., Jinggui F., Tomer E., Hillel J., Lavi U Cultivar identification and genetic map of mango (Mangifera indica). EUPHYTICA 122(1): Gurevitz S., Lavi U., Cohen Y Genetic variation in date palms propagated from offshoots and tissue culture. J. AMER. SOC. HORT. SCI. (in press). Vos P., Hogers R., Bleeker M., Rejans M., van de Lee T., Homes M., Frijters A., Pot J., Peleman J., Kuiper M., Zabeau M AFLP: a new technique for DNA fingerprinting. NUC. ACID. RES. 23: J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006: 13-19
7 Potential applications of modern biological tech niques in breeding MOŻLIWOŚCI ZASTOSOWANIA TECHNIK NOWOCZESNEJ BIOLOGII W HODOWLI DRZEW OWOCOWYCH Uri Lavi, Slava Gurevitz, Giara Ben Ari, David Saada, Khalib Kashkush, Tali Paz, Te Twito, Yuval Cohen, Jossi Hillel i Giora Simchen S T R E S Z C Z E N I E Klasyczna hodowla roślin drzewiastych ma wiele ograniczeń. Długi okres juwenilny i wielkośćroślin powoduje, że tradycyjne metody oparte na długotrwałej selekcji są czasochłonne i kosztowne. Obniżenie kosztów hodowli i jej przyspieszenie jest możliwe poprzez zastosowanie metod biotechnologicznych. Jednąz nowoczesnych strategii jest inżynieria genetyczna, umożliwiająca wprowadzenie genów kodujących pożądane cechy do genomu roślin. Technika ta nie dysponuje jednak wystarczająco wydajnymi metodykami transformacji i regeneracji roślin drzewiastych. Ograniczona jest również dostępnośćpotencjalnych transgenów, które mogłyby sterowaćzłożonymi procesami. W przeciwieństwie do strategii wykorzystujących inżynierięgenetycznątechniki oparte lub/1 związane z analizągenomu (różnicowanie roślin na podstawie ich wzorów genetycznych, identyfikacja ważnych genów, selekcja oparta na markerach molekularnych MAS, zastosowanie mikromacierzy i technik bioinformatycznych), sąjuż dokładnie opracowane. Przydatnośćtych technik dla hodowli roślin drzewiastych została wykazana w artykule w oparciu o szczegółowe badania przeprowadzone w Volcani Centre. Słowa kluczowe: markery AFLP, SNP, introgresja genów, bioróżnorodność J. Fruit Ornam. Plant Res. vol. 14 (Suppl. 1), 2006:
Tychy, plan miasta: Skala 1: (Polish Edition)
 Tychy, plan miasta: Skala 1:20 000 (Polish Edition) Poland) Przedsiebiorstwo Geodezyjno-Kartograficzne (Katowice Click here if your download doesn"t start automatically Tychy, plan miasta: Skala 1:20 000
Tychy, plan miasta: Skala 1:20 000 (Polish Edition) Poland) Przedsiebiorstwo Geodezyjno-Kartograficzne (Katowice Click here if your download doesn"t start automatically Tychy, plan miasta: Skala 1:20 000
Zakopane, plan miasta: Skala ok. 1: = City map (Polish Edition)
 Zakopane, plan miasta: Skala ok. 1:15 000 = City map (Polish Edition) Click here if your download doesn"t start automatically Zakopane, plan miasta: Skala ok. 1:15 000 = City map (Polish Edition) Zakopane,
Zakopane, plan miasta: Skala ok. 1:15 000 = City map (Polish Edition) Click here if your download doesn"t start automatically Zakopane, plan miasta: Skala ok. 1:15 000 = City map (Polish Edition) Zakopane,
SSW1.1, HFW Fry #20, Zeno #25 Benchmark: Qtr.1. Fry #65, Zeno #67. like
 SSW1.1, HFW Fry #20, Zeno #25 Benchmark: Qtr.1 I SSW1.1, HFW Fry #65, Zeno #67 Benchmark: Qtr.1 like SSW1.2, HFW Fry #47, Zeno #59 Benchmark: Qtr.1 do SSW1.2, HFW Fry #5, Zeno #4 Benchmark: Qtr.1 to SSW1.2,
SSW1.1, HFW Fry #20, Zeno #25 Benchmark: Qtr.1 I SSW1.1, HFW Fry #65, Zeno #67 Benchmark: Qtr.1 like SSW1.2, HFW Fry #47, Zeno #59 Benchmark: Qtr.1 do SSW1.2, HFW Fry #5, Zeno #4 Benchmark: Qtr.1 to SSW1.2,
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition)
 Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Patients price acceptance SELECTED FINDINGS
 Patients price acceptance SELECTED FINDINGS October 2015 Summary With growing economy and Poles benefiting from this growth, perception of prices changes - this is also true for pharmaceuticals It may
Patients price acceptance SELECTED FINDINGS October 2015 Summary With growing economy and Poles benefiting from this growth, perception of prices changes - this is also true for pharmaceuticals It may
Karpacz, plan miasta 1:10 000: Panorama Karkonoszy, mapa szlakow turystycznych (Polish Edition)
 Karpacz, plan miasta 1:10 000: Panorama Karkonoszy, mapa szlakow turystycznych (Polish Edition) J Krupski Click here if your download doesn"t start automatically Karpacz, plan miasta 1:10 000: Panorama
Karpacz, plan miasta 1:10 000: Panorama Karkonoszy, mapa szlakow turystycznych (Polish Edition) J Krupski Click here if your download doesn"t start automatically Karpacz, plan miasta 1:10 000: Panorama
ARNOLD. EDUKACJA KULTURYSTY (POLSKA WERSJA JEZYKOWA) BY DOUGLAS KENT HALL
 Read Online and Download Ebook ARNOLD. EDUKACJA KULTURYSTY (POLSKA WERSJA JEZYKOWA) BY DOUGLAS KENT HALL DOWNLOAD EBOOK : ARNOLD. EDUKACJA KULTURYSTY (POLSKA WERSJA Click link bellow and free register
Read Online and Download Ebook ARNOLD. EDUKACJA KULTURYSTY (POLSKA WERSJA JEZYKOWA) BY DOUGLAS KENT HALL DOWNLOAD EBOOK : ARNOLD. EDUKACJA KULTURYSTY (POLSKA WERSJA Click link bellow and free register
ERASMUS + : Trail of extinct and active volcanoes, earthquakes through Europe. SURVEY TO STUDENTS.
 ERASMUS + : Trail of extinct and active volcanoes, earthquakes through Europe. SURVEY TO STUDENTS. Strona 1 1. Please give one answer. I am: Students involved in project 69% 18 Student not involved in
ERASMUS + : Trail of extinct and active volcanoes, earthquakes through Europe. SURVEY TO STUDENTS. Strona 1 1. Please give one answer. I am: Students involved in project 69% 18 Student not involved in
Cracow University of Economics Poland. Overview. Sources of Real GDP per Capita Growth: Polish Regional-Macroeconomic Dimensions 2000-2005
 Cracow University of Economics Sources of Real GDP per Capita Growth: Polish Regional-Macroeconomic Dimensions 2000-2005 - Key Note Speech - Presented by: Dr. David Clowes The Growth Research Unit CE Europe
Cracow University of Economics Sources of Real GDP per Capita Growth: Polish Regional-Macroeconomic Dimensions 2000-2005 - Key Note Speech - Presented by: Dr. David Clowes The Growth Research Unit CE Europe
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition)
 Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Machine Learning for Data Science (CS4786) Lecture11. Random Projections & Canonical Correlation Analysis
 Machine Learning for Data Science (CS4786) Lecture11 5 Random Projections & Canonical Correlation Analysis The Tall, THE FAT AND THE UGLY n X d The Tall, THE FAT AND THE UGLY d X > n X d n = n d d The
Machine Learning for Data Science (CS4786) Lecture11 5 Random Projections & Canonical Correlation Analysis The Tall, THE FAT AND THE UGLY n X d The Tall, THE FAT AND THE UGLY d X > n X d n = n d d The
Network Services for Spatial Data in European Geo-Portals and their Compliance with ISO and OGC Standards
 INSPIRE Conference 2010 INSPIRE as a Framework for Cooperation Network Services for Spatial Data in European Geo-Portals and their Compliance with ISO and OGC Standards Elżbieta Bielecka Agnieszka Zwirowicz
INSPIRE Conference 2010 INSPIRE as a Framework for Cooperation Network Services for Spatial Data in European Geo-Portals and their Compliance with ISO and OGC Standards Elżbieta Bielecka Agnieszka Zwirowicz
 www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
OpenPoland.net API Documentation
 OpenPoland.net API Documentation Release 1.0 Michał Gryczka July 11, 2014 Contents 1 REST API tokens: 3 1.1 How to get a token............................................ 3 2 REST API : search for assets
OpenPoland.net API Documentation Release 1.0 Michał Gryczka July 11, 2014 Contents 1 REST API tokens: 3 1.1 How to get a token............................................ 3 2 REST API : search for assets
Emilka szuka swojej gwiazdy / Emily Climbs (Emily, #2)
 Emilka szuka swojej gwiazdy / Emily Climbs (Emily, #2) Click here if your download doesn"t start automatically Emilka szuka swojej gwiazdy / Emily Climbs (Emily, #2) Emilka szuka swojej gwiazdy / Emily
Emilka szuka swojej gwiazdy / Emily Climbs (Emily, #2) Click here if your download doesn"t start automatically Emilka szuka swojej gwiazdy / Emily Climbs (Emily, #2) Emilka szuka swojej gwiazdy / Emily
MaPlan Sp. z O.O. Click here if your download doesn"t start automatically
 Mierzeja Wislana, mapa turystyczna 1:50 000: Mikoszewo, Jantar, Stegna, Sztutowo, Katy Rybackie, Przebrno, Krynica Morska, Piaski, Frombork =... = Carte touristique (Polish Edition) MaPlan Sp. z O.O Click
Mierzeja Wislana, mapa turystyczna 1:50 000: Mikoszewo, Jantar, Stegna, Sztutowo, Katy Rybackie, Przebrno, Krynica Morska, Piaski, Frombork =... = Carte touristique (Polish Edition) MaPlan Sp. z O.O Click
SubVersion. Piotr Mikulski. SubVersion. P. Mikulski. Co to jest subversion? Zalety SubVersion. Wady SubVersion. Inne różnice SubVersion i CVS
 Piotr Mikulski 2006 Subversion is a free/open-source version control system. That is, Subversion manages files and directories over time. A tree of files is placed into a central repository. The repository
Piotr Mikulski 2006 Subversion is a free/open-source version control system. That is, Subversion manages files and directories over time. A tree of files is placed into a central repository. The repository
Revenue Maximization. Sept. 25, 2018
 Revenue Maximization Sept. 25, 2018 Goal So Far: Ideal Auctions Dominant-Strategy Incentive Compatible (DSIC) b i = v i is a dominant strategy u i 0 x is welfare-maximizing x and p run in polynomial time
Revenue Maximization Sept. 25, 2018 Goal So Far: Ideal Auctions Dominant-Strategy Incentive Compatible (DSIC) b i = v i is a dominant strategy u i 0 x is welfare-maximizing x and p run in polynomial time
 www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
Helena Boguta, klasa 8W, rok szkolny 2018/2019
 Poniższy zbiór zadań został wykonany w ramach projektu Mazowiecki program stypendialny dla uczniów szczególnie uzdolnionych - najlepsza inwestycja w człowieka w roku szkolnym 2018/2019. Składają się na
Poniższy zbiór zadań został wykonany w ramach projektu Mazowiecki program stypendialny dla uczniów szczególnie uzdolnionych - najlepsza inwestycja w człowieka w roku szkolnym 2018/2019. Składają się na
Stargard Szczecinski i okolice (Polish Edition)
 Stargard Szczecinski i okolice (Polish Edition) Janusz Leszek Jurkiewicz Click here if your download doesn"t start automatically Stargard Szczecinski i okolice (Polish Edition) Janusz Leszek Jurkiewicz
Stargard Szczecinski i okolice (Polish Edition) Janusz Leszek Jurkiewicz Click here if your download doesn"t start automatically Stargard Szczecinski i okolice (Polish Edition) Janusz Leszek Jurkiewicz
Katowice, plan miasta: Skala 1: = City map = Stadtplan (Polish Edition)
 Katowice, plan miasta: Skala 1:20 000 = City map = Stadtplan (Polish Edition) Polskie Przedsiebiorstwo Wydawnictw Kartograficznych im. Eugeniusza Romera Click here if your download doesn"t start automatically
Katowice, plan miasta: Skala 1:20 000 = City map = Stadtplan (Polish Edition) Polskie Przedsiebiorstwo Wydawnictw Kartograficznych im. Eugeniusza Romera Click here if your download doesn"t start automatically
Domy inaczej pomyślane A different type of housing CEZARY SANKOWSKI
 Domy inaczej pomyślane A different type of housing CEZARY SANKOWSKI O tym, dlaczego warto budować pasywnie, komu budownictwo pasywne się opłaca, a kto się go boi, z architektem, Cezarym Sankowskim, rozmawia
Domy inaczej pomyślane A different type of housing CEZARY SANKOWSKI O tym, dlaczego warto budować pasywnie, komu budownictwo pasywne się opłaca, a kto się go boi, z architektem, Cezarym Sankowskim, rozmawia
 aforementioned device she also has to estimate the time when the patients need the infusion to be replaced and/or disconnected. Meanwhile, however, she must cope with many other tasks. If the department
aforementioned device she also has to estimate the time when the patients need the infusion to be replaced and/or disconnected. Meanwhile, however, she must cope with many other tasks. If the department
Rozpoznawanie twarzy metodą PCA Michał Bereta 1. Testowanie statystycznej istotności różnic między jakością klasyfikatorów
 Rozpoznawanie twarzy metodą PCA Michał Bereta www.michalbereta.pl 1. Testowanie statystycznej istotności różnic między jakością klasyfikatorów Wiemy, że możemy porównywad klasyfikatory np. za pomocą kroswalidacji.
Rozpoznawanie twarzy metodą PCA Michał Bereta www.michalbereta.pl 1. Testowanie statystycznej istotności różnic między jakością klasyfikatorów Wiemy, że możemy porównywad klasyfikatory np. za pomocą kroswalidacji.
Projekty Marie Curie Actions w praktyce: EGALITE (IAPP) i ArSInformatiCa (IOF)
 Gliwice, Poland, 28th February 2014 Projekty Marie Curie Actions w praktyce: EGALITE (IAPP) i ArSInformatiCa (IOF) Krzysztof A. Cyran The project has received Community research funding under the 7th Framework
Gliwice, Poland, 28th February 2014 Projekty Marie Curie Actions w praktyce: EGALITE (IAPP) i ArSInformatiCa (IOF) Krzysztof A. Cyran The project has received Community research funding under the 7th Framework
Machine Learning for Data Science (CS4786) Lecture 11. Spectral Embedding + Clustering
 Machine Learning for Data Science (CS4786) Lecture 11 Spectral Embedding + Clustering MOTIVATING EXAMPLE What can you say from this network? MOTIVATING EXAMPLE How about now? THOUGHT EXPERIMENT For each
Machine Learning for Data Science (CS4786) Lecture 11 Spectral Embedding + Clustering MOTIVATING EXAMPLE What can you say from this network? MOTIVATING EXAMPLE How about now? THOUGHT EXPERIMENT For each
Bioinformatyka wykład I.2009
 Bioinformatyka wykład 13 20.I.2009 biologia systemów biologiczne dane wielowymiarowe Krzysztof Pawłowski Krzysztof_Pawlowski@sggw.pl 2009-01-22 1 Plan wykładu Biologia systemów Bazy danych ekspresji genów
Bioinformatyka wykład 13 20.I.2009 biologia systemów biologiczne dane wielowymiarowe Krzysztof Pawłowski Krzysztof_Pawlowski@sggw.pl 2009-01-22 1 Plan wykładu Biologia systemów Bazy danych ekspresji genów
DOI: / /32/37
 . 2015. 4 (32) 1:18 DOI: 10.17223/1998863 /32/37 -,,. - -. :,,,,., -, -.,.-.,.,.,. -., -,.,,., -, 70 80. (.,.,. ),, -,.,, -,, (1886 1980).,.,, (.,.,..), -, -,,,, ; -, - 346, -,.. :, -, -,,,,,.,,, -,,,
. 2015. 4 (32) 1:18 DOI: 10.17223/1998863 /32/37 -,,. - -. :,,,,., -, -.,.-.,.,.,. -., -,.,,., -, 70 80. (.,.,. ),, -,.,, -,, (1886 1980).,.,, (.,.,..), -, -,,,, ; -, - 346, -,.. :, -, -,,,,,.,,, -,,,
EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO
 Miejsce na naklejkę z kodem szkoły dysleksja MJA-R1_1P-072 EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO MAJ ROK 2007 Instrukcja dla zdającego POZIOM ROZSZERZONY CZĘŚĆ I Czas pracy 120 minut 1. Sprawdź, czy
Miejsce na naklejkę z kodem szkoły dysleksja MJA-R1_1P-072 EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO MAJ ROK 2007 Instrukcja dla zdającego POZIOM ROZSZERZONY CZĘŚĆ I Czas pracy 120 minut 1. Sprawdź, czy
European Crime Prevention Award (ECPA) Annex I - new version 2014
 European Crime Prevention Award (ECPA) Annex I - new version 2014 Załącznik nr 1 General information (Informacje ogólne) 1. Please specify your country. (Kraj pochodzenia:) 2. Is this your country s ECPA
European Crime Prevention Award (ECPA) Annex I - new version 2014 Załącznik nr 1 General information (Informacje ogólne) 1. Please specify your country. (Kraj pochodzenia:) 2. Is this your country s ECPA
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition)
 Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Zarządzanie sieciami telekomunikacyjnymi
 SNMP Protocol The Simple Network Management Protocol (SNMP) is an application layer protocol that facilitates the exchange of management information between network devices. It is part of the Transmission
SNMP Protocol The Simple Network Management Protocol (SNMP) is an application layer protocol that facilitates the exchange of management information between network devices. It is part of the Transmission
Analysis of Movie Profitability STAT 469 IN CLASS ANALYSIS #2
 Analysis of Movie Profitability STAT 469 IN CLASS ANALYSIS #2 aaaklnictzzjb9tgfmcnadpg7oy0lxa9edva9kkapdarhyk2k7gourinlwsweyzikuyiigvyleiv/cv767fpf/5crc1xt9va5mx7w3m/ecuqw1kuztpx/rl3/70h73/w4cog9dhhn3z62d6jzy+yzj766txpoir9nzszisjynetqr+rvlfvyoozu5xbybpsxb1wahul8phczdt2v4zgchb7uecwphlyigrgkjcyiflfyci0kxnmr4z6kw0jsokvot8isntpa3gbknlcufiv/h+hh+eur4fomd417rvtfjoit5pfju6yxiab2fmwk0y/feuybobqk+axnke8xzjjhfyd8kkpl9zdoddkazd5j6bzpemjb64smjb6vb4xmehysu08lsrszopxftlzee130jcb0zjxy7r5wa2f1s2off2+dyatrughnrtpkuprlcpu55zlxpss/yqe2eamjkcf0jye8w8yas0paf6t0t2i9stmcua+inbi2rt01tz22tubbqwidypvgz6piynkpobirkxgu54ibzoti4pkw2i5ow9lnuaoabhuxfxqhvnrj6w15tb3furnbm+scyxobjhr5pmj5j/w5ix9wsa2tlwx9alpshlunzjgnrwvqbpwzjl9wes+ptyn+ypy/jgskavtl8j0hz1djdhzwtpjbbvpr1zj7jpg6ve7zxfngj75zee0vmp9qm2uvgu/9zdofq6r+g8l4xctvo+v+xdrfr8oxiwutycu0qgyf8icuyvp/sixfi9zxe11vp6mrjjovpmxm6acrtbia+wjr9bevlgjwlz5xd3rfna9g06qytaoofk8olxbxc7xby2evqjmmk6pjvvzxmpbnct6+036xp5vdbrnbdqph8brlfn/n/khnfumhf6z1v7h/80yieukkd5j0un82t9mynxzmk0s/bzn4tacdziszdhwrl8x5ako8qp1n1zn0k6w2em0km9zj1i4yt1pt3xiprw85jmc2m1ut2geum6y6es2fwx6c+wlrpykblopbuj5nnr2byygfy5opllv4+jmm7s6u+tvhywbnb0kv2lt5th4xipmiij+y1toiyo7bo0d+vzvovjkp6aoejsubhj3qrp3fjd/m23pay8h218ibvx3nicofvd1xi86+kh6nb/b+hgsjp5+qwpurzlir15np66vmdehh6tyazdm1k/5ejtuvurgcqux6yc+qw/sbsaj7lkt4x9qmtp7euk6zbdedyuzu6ptsu2eeu3rxcz06uf6g8wyuveznhkbzynajbb7r7cbmla+jbtrst0ow2v6ntkwv8svnwqnu5pa3oxfeexf93739p93chq/fv+jr8r0d9brhpcxr2w88bvqbr41j6wvrb+u5dzjpvx+veoaxwptzp/8cen+xbg==
Analysis of Movie Profitability STAT 469 IN CLASS ANALYSIS #2 aaaklnictzzjb9tgfmcnadpg7oy0lxa9edva9kkapdarhyk2k7gourinlwsweyzikuyiigvyleiv/cv767fpf/5crc1xt9va5mx7w3m/ecuqw1kuztpx/rl3/70h73/w4cog9dhhn3z62d6jzy+yzj766txpoir9nzszisjynetqr+rvlfvyoozu5xbybpsxb1wahul8phczdt2v4zgchb7uecwphlyigrgkjcyiflfyci0kxnmr4z6kw0jsokvot8isntpa3gbknlcufiv/h+hh+eur4fomd417rvtfjoit5pfju6yxiab2fmwk0y/feuybobqk+axnke8xzjjhfyd8kkpl9zdoddkazd5j6bzpemjb64smjb6vb4xmehysu08lsrszopxftlzee130jcb0zjxy7r5wa2f1s2off2+dyatrughnrtpkuprlcpu55zlxpss/yqe2eamjkcf0jye8w8yas0paf6t0t2i9stmcua+inbi2rt01tz22tubbqwidypvgz6piynkpobirkxgu54ibzoti4pkw2i5ow9lnuaoabhuxfxqhvnrj6w15tb3furnbm+scyxobjhr5pmj5j/w5ix9wsa2tlwx9alpshlunzjgnrwvqbpwzjl9wes+ptyn+ypy/jgskavtl8j0hz1djdhzwtpjbbvpr1zj7jpg6ve7zxfngj75zee0vmp9qm2uvgu/9zdofq6r+g8l4xctvo+v+xdrfr8oxiwutycu0qgyf8icuyvp/sixfi9zxe11vp6mrjjovpmxm6acrtbia+wjr9bevlgjwlz5xd3rfna9g06qytaoofk8olxbxc7xby2evqjmmk6pjvvzxmpbnct6+036xp5vdbrnbdqph8brlfn/n/khnfumhf6z1v7h/80yieukkd5j0un82t9mynxzmk0s/bzn4tacdziszdhwrl8x5ako8qp1n1zn0k6w2em0km9zj1i4yt1pt3xiprw85jmc2m1ut2geum6y6es2fwx6c+wlrpykblopbuj5nnr2byygfy5opllv4+jmm7s6u+tvhywbnb0kv2lt5th4xipmiij+y1toiyo7bo0d+vzvovjkp6aoejsubhj3qrp3fjd/m23pay8h218ibvx3nicofvd1xi86+kh6nb/b+hgsjp5+qwpurzlir15np66vmdehh6tyazdm1k/5ejtuvurgcqux6yc+qw/sbsaj7lkt4x9qmtp7euk6zbdedyuzu6ptsu2eeu3rxcz06uf6g8wyuveznhkbzynajbb7r7cbmla+jbtrst0ow2v6ntkwv8svnwqnu5pa3oxfeexf93739p93chq/fv+jr8r0d9brhpcxr2w88bvqbr41j6wvrb+u5dzjpvx+veoaxwptzp/8cen+xbg==
The Overview of Civilian Applications of Airborne SAR Systems
 The Overview of Civilian Applications of Airborne SAR Systems Maciej Smolarczyk, Piotr Samczyński Andrzej Gadoś, Maj Mordzonek Research and Development Department of PIT S.A. PART I WHAT DOES SAR MEAN?
The Overview of Civilian Applications of Airborne SAR Systems Maciej Smolarczyk, Piotr Samczyński Andrzej Gadoś, Maj Mordzonek Research and Development Department of PIT S.A. PART I WHAT DOES SAR MEAN?
Proposal of thesis topic for mgr in. (MSE) programme in Telecommunications and Computer Science
 Proposal of thesis topic for mgr in (MSE) programme 1 Topic: Monte Carlo Method used for a prognosis of a selected technological process 2 Supervisor: Dr in Małgorzata Langer 3 Auxiliary supervisor: 4
Proposal of thesis topic for mgr in (MSE) programme 1 Topic: Monte Carlo Method used for a prognosis of a selected technological process 2 Supervisor: Dr in Małgorzata Langer 3 Auxiliary supervisor: 4
Jak zasada Pareto może pomóc Ci w nauce języków obcych?
 Jak zasada Pareto może pomóc Ci w nauce języków obcych? Artykuł pobrano ze strony eioba.pl Pokazuje, jak zastosowanie zasady Pareto może usprawnić Twoją naukę angielskiego. Słynna zasada Pareto mówi o
Jak zasada Pareto może pomóc Ci w nauce języków obcych? Artykuł pobrano ze strony eioba.pl Pokazuje, jak zastosowanie zasady Pareto może usprawnić Twoją naukę angielskiego. Słynna zasada Pareto mówi o
Cracow University of Economics Poland
 Cracow University of Economics Poland Sources of Real GDP per Capita Growth: Polish Regional-Macroeconomic Dimensions 2000-2005 - Keynote Speech - Presented by: Dr. David Clowes The Growth Research Unit,
Cracow University of Economics Poland Sources of Real GDP per Capita Growth: Polish Regional-Macroeconomic Dimensions 2000-2005 - Keynote Speech - Presented by: Dr. David Clowes The Growth Research Unit,
Pielgrzymka do Ojczyzny: Przemowienia i homilie Ojca Swietego Jana Pawla II (Jan Pawel II-- pierwszy Polak na Stolicy Piotrowej) (Polish Edition)
 Pielgrzymka do Ojczyzny: Przemowienia i homilie Ojca Swietego Jana Pawla II (Jan Pawel II-- pierwszy Polak na Stolicy Piotrowej) (Polish Edition) Click here if your download doesn"t start automatically
Pielgrzymka do Ojczyzny: Przemowienia i homilie Ojca Swietego Jana Pawla II (Jan Pawel II-- pierwszy Polak na Stolicy Piotrowej) (Polish Edition) Click here if your download doesn"t start automatically
Krytyczne czynniki sukcesu w zarządzaniu projektami
 Seweryn SPAŁEK Krytyczne czynniki sukcesu w zarządzaniu projektami MONOGRAFIA Wydawnictwo Politechniki Śląskiej Gliwice 2004 SPIS TREŚCI WPROWADZENIE 5 1. ZARZĄDZANIE PROJEKTAMI W ORGANIZACJI 13 1.1. Zarządzanie
Seweryn SPAŁEK Krytyczne czynniki sukcesu w zarządzaniu projektami MONOGRAFIA Wydawnictwo Politechniki Śląskiej Gliwice 2004 SPIS TREŚCI WPROWADZENIE 5 1. ZARZĄDZANIE PROJEKTAMI W ORGANIZACJI 13 1.1. Zarządzanie
EXAMPLES OF CABRI GEOMETRE II APPLICATION IN GEOMETRIC SCIENTIFIC RESEARCH
 Anna BŁACH Centre of Geometry and Engineering Graphics Silesian University of Technology in Gliwice EXAMPLES OF CABRI GEOMETRE II APPLICATION IN GEOMETRIC SCIENTIFIC RESEARCH Introduction Computer techniques
Anna BŁACH Centre of Geometry and Engineering Graphics Silesian University of Technology in Gliwice EXAMPLES OF CABRI GEOMETRE II APPLICATION IN GEOMETRIC SCIENTIFIC RESEARCH Introduction Computer techniques
Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition)
 Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition) Piotr Maluskiewicz Click here if your download doesn"t start automatically Miedzy
Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition) Piotr Maluskiewicz Click here if your download doesn"t start automatically Miedzy
Latent Dirichlet Allocation Models and their Evaluation IT for Practice 2016
 Latent Dirichlet Allocation Models and their Evaluation IT for Practice 2016 Paweł Lula Cracow University of Economics, Poland pawel.lula@uek.krakow.pl Latent Dirichlet Allocation (LDA) Documents Latent
Latent Dirichlet Allocation Models and their Evaluation IT for Practice 2016 Paweł Lula Cracow University of Economics, Poland pawel.lula@uek.krakow.pl Latent Dirichlet Allocation (LDA) Documents Latent
Pro-tumoral immune cell alterations in wild type and Shbdeficient mice in response to 4T1 breast carcinomas
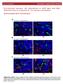 www.oncotarget.com Oncotarget, Supplementary Materials Pro-tumoral immune cell alterations in wild type and Shbdeficient mice in response to 4T1 breast carcinomas SUPPLEMENTARY MATERIALS Supplementary
www.oncotarget.com Oncotarget, Supplementary Materials Pro-tumoral immune cell alterations in wild type and Shbdeficient mice in response to 4T1 breast carcinomas SUPPLEMENTARY MATERIALS Supplementary
Fig 5 Spectrograms of the original signal (top) extracted shaft-related GAD components (middle) and
 Fig 4 Measured vibration signal (top). Blue original signal. Red component related to periodic excitation of resonances and noise. Green component related. Rotational speed profile used for experiment
Fig 4 Measured vibration signal (top). Blue original signal. Red component related to periodic excitation of resonances and noise. Green component related. Rotational speed profile used for experiment
Weronika Mysliwiec, klasa 8W, rok szkolny 2018/2019
 Poniższy zbiór zadań został wykonany w ramach projektu Mazowiecki program stypendialny dla uczniów szczególnie uzdolnionych - najlepsza inwestycja w człowieka w roku szkolnym 2018/2019. Tresci zadań rozwiązanych
Poniższy zbiór zadań został wykonany w ramach projektu Mazowiecki program stypendialny dla uczniów szczególnie uzdolnionych - najlepsza inwestycja w człowieka w roku szkolnym 2018/2019. Tresci zadań rozwiązanych
ABOUT NEW EASTERN EUROPE BESTmQUARTERLYmJOURNAL
 ABOUT NEW EASTERN EUROPE BESTmQUARTERLYmJOURNAL Formanminsidemlookmatmpoliticsxmculturexmsocietymandm economyminmthemregionmofmcentralmandmeasternm EuropexmtheremismnomothermsourcemlikemNew Eastern EuropeImSincemitsmlaunchminmPw--xmthemmagazinemhasm
ABOUT NEW EASTERN EUROPE BESTmQUARTERLYmJOURNAL Formanminsidemlookmatmpoliticsxmculturexmsocietymandm economyminmthemregionmofmcentralmandmeasternm EuropexmtheremismnomothermsourcemlikemNew Eastern EuropeImSincemitsmlaunchminmPw--xmthemmagazinemhasm
Ankiety Nowe funkcje! Pomoc magda.szewczyk@slo-wroc.pl. magda.szewczyk@slo-wroc.pl. Twoje konto Wyloguj. BIODIVERSITY OF RIVERS: Survey to students
 Ankiety Nowe funkcje! Pomoc magda.szewczyk@slo-wroc.pl Back Twoje konto Wyloguj magda.szewczyk@slo-wroc.pl BIODIVERSITY OF RIVERS: Survey to students Tworzenie ankiety Udostępnianie Analiza (55) Wyniki
Ankiety Nowe funkcje! Pomoc magda.szewczyk@slo-wroc.pl Back Twoje konto Wyloguj magda.szewczyk@slo-wroc.pl BIODIVERSITY OF RIVERS: Survey to students Tworzenie ankiety Udostępnianie Analiza (55) Wyniki
Working Tax Credit Child Tax Credit Jobseeker s Allowance
 Benefits Depending on your residency status (EU citizen or not) there are various benefits available to help you with costs of living. A8 nationals need to have been working for a year and be registered
Benefits Depending on your residency status (EU citizen or not) there are various benefits available to help you with costs of living. A8 nationals need to have been working for a year and be registered
Few-fermion thermometry
 Few-fermion thermometry Phys. Rev. A 97, 063619 (2018) Tomasz Sowiński Institute of Physics of the Polish Academy of Sciences Co-authors: Marcin Płodzień Rafał Demkowicz-Dobrzański FEW-BODY PROBLEMS FewBody.ifpan.edu.pl
Few-fermion thermometry Phys. Rev. A 97, 063619 (2018) Tomasz Sowiński Institute of Physics of the Polish Academy of Sciences Co-authors: Marcin Płodzień Rafał Demkowicz-Dobrzański FEW-BODY PROBLEMS FewBody.ifpan.edu.pl
Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition)
 Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition) Piotr Maluskiewicz Click here if your download doesn"t start automatically Miedzy
Miedzy legenda a historia: Szlakiem piastowskim z Poznania do Gniezna (Biblioteka Kroniki Wielkopolski) (Polish Edition) Piotr Maluskiewicz Click here if your download doesn"t start automatically Miedzy
Sargent Opens Sonairte Farmers' Market
 Sargent Opens Sonairte Farmers' Market 31 March, 2008 1V8VIZSV7EVKIRX8(1MRMWXIVSJ7XEXIEXXLI(ITEVXQIRXSJ%KVMGYPXYVI *MWLIVMIWERH*SSHTIVJSVQIHXLISJJMGMEPSTIRMRKSJXLI7SREMVXI*EVQIVW 1EVOIXMR0E]XS[R'S1IEXL
Sargent Opens Sonairte Farmers' Market 31 March, 2008 1V8VIZSV7EVKIRX8(1MRMWXIVSJ7XEXIEXXLI(ITEVXQIRXSJ%KVMGYPXYVI *MWLIVMIWERH*SSHTIVJSVQIHXLISJJMGMEPSTIRMRKSJXLI7SREMVXI*EVQIVW 1EVOIXMR0E]XS[R'S1IEXL
Wyniki Finansowe Grupy Kapitałowej Private Equity Managers S.A. za Warszawa, 29 Marca 2017
 Wyniki Finansowe Grupy Kapitałowej Private Equity Managers S.A. za 2016 Warszawa, 29 Marca 2017 01 wyniki finansowe GK PEM 01 wyniki finansowe GK PEM W 2016 roku GK PEM zrealizowała PLN 14,4 mln wyniku
Wyniki Finansowe Grupy Kapitałowej Private Equity Managers S.A. za 2016 Warszawa, 29 Marca 2017 01 wyniki finansowe GK PEM 01 wyniki finansowe GK PEM W 2016 roku GK PEM zrealizowała PLN 14,4 mln wyniku
Egzamin maturalny z języka angielskiego na poziomie dwujęzycznym Rozmowa wstępna (wyłącznie dla egzaminującego)
 112 Informator o egzaminie maturalnym z języka angielskiego od roku szkolnego 2014/2015 2.6.4. Część ustna. Przykładowe zestawy zadań Przykładowe pytania do rozmowy wstępnej Rozmowa wstępna (wyłącznie
112 Informator o egzaminie maturalnym z języka angielskiego od roku szkolnego 2014/2015 2.6.4. Część ustna. Przykładowe zestawy zadań Przykładowe pytania do rozmowy wstępnej Rozmowa wstępna (wyłącznie
Hard-Margin Support Vector Machines
 Hard-Margin Support Vector Machines aaacaxicbzdlssnafiyn9vbjlepk3ay2gicupasvu4iblxuaw2hjmuwn7ddjjmxm1bkcg1/fjqsvt76fo9/gazqfvn8y+pjpozw5vx8zkpvtfxmlhcwl5zxyqrm2vrg5zw3vxmsoezi4ogkr6phieky5crvvjhriqvdom9l2xxftevuwcekj3lktmhghgniauiyutvrwxtvme34a77kbvg73gtygpjsrfati1+xc8c84bvraowbf+uwnipyehcvmkjrdx46vlykhkgykm3ujjdhcyzqkxy0chur6ax5cbg+1m4bbjptjcubuz4kuhvjoql93hkin5hxtav5x6yyqopnsyuneey5ni4keqrxbar5wqaxbik00icyo/iveiyqqvjo1u4fgzj/8f9x67bzmxnurjzmijtlybwfgcdjgfdtajwgcf2dwaj7ac3g1ho1n4814n7wwjgjmf/ys8fenfycuzq==
Hard-Margin Support Vector Machines aaacaxicbzdlssnafiyn9vbjlepk3ay2gicupasvu4iblxuaw2hjmuwn7ddjjmxm1bkcg1/fjqsvt76fo9/gazqfvn8y+pjpozw5vx8zkpvtfxmlhcwl5zxyqrm2vrg5zw3vxmsoezi4ogkr6phieky5crvvjhriqvdom9l2xxftevuwcekj3lktmhghgniauiyutvrwxtvme34a77kbvg73gtygpjsrfati1+xc8c84bvraowbf+uwnipyehcvmkjrdx46vlykhkgykm3ujjdhcyzqkxy0chur6ax5cbg+1m4bbjptjcubuz4kuhvjoql93hkin5hxtav5x6yyqopnsyuneey5ni4keqrxbar5wqaxbik00icyo/iveiyqqvjo1u4fgzj/8f9x67bzmxnurjzmijtlybwfgcdjgfdtajwgcf2dwaj7ac3g1ho1n4814n7wwjgjmf/ys8fenfycuzq==
 www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
deep learning for NLP (5 lectures)
 TTIC 31210: Advanced Natural Language Processing Kevin Gimpel Spring 2019 Lecture 6: Finish Transformers; Sequence- to- Sequence Modeling and AJenKon 1 Roadmap intro (1 lecture) deep learning for NLP (5
TTIC 31210: Advanced Natural Language Processing Kevin Gimpel Spring 2019 Lecture 6: Finish Transformers; Sequence- to- Sequence Modeling and AJenKon 1 Roadmap intro (1 lecture) deep learning for NLP (5
Baptist Church Records
 Baptist Church Records The Baptist religion was a religious minority in Poland, making it more difficult to know when and where records of this religion might be available. In an article from Rodziny,
Baptist Church Records The Baptist religion was a religious minority in Poland, making it more difficult to know when and where records of this religion might be available. In an article from Rodziny,
Blow-Up: Photographs in the Time of Tumult; Black and White Photography Festival Zakopane Warszawa 2002 / Powiekszenie: Fotografie w czasach zgielku
 Blow-Up: Photographs in the Time of Tumult; Black and White Photography Festival Zakopane Warszawa 2002 / Powiekszenie: Fotografie w czasach zgielku Juliusz and Maciej Zalewski eds. and A. D. Coleman et
Blow-Up: Photographs in the Time of Tumult; Black and White Photography Festival Zakopane Warszawa 2002 / Powiekszenie: Fotografie w czasach zgielku Juliusz and Maciej Zalewski eds. and A. D. Coleman et
What our clients think about us? A summary od survey results
 What our clients think about us? A summary od survey results customer satisfaction survey We conducted our audit in June 2015 This is the first survey about customer satisfaction Why? To get customer feedback
What our clients think about us? A summary od survey results customer satisfaction survey We conducted our audit in June 2015 This is the first survey about customer satisfaction Why? To get customer feedback
Probability definition
 Probability Probability definition Probability of an event P(A) denotes the frequency of it s occurrence in a long (infinite) series of repeats P( A) = A Ω Ω A 0 P It s s represented by a number in the
Probability Probability definition Probability of an event P(A) denotes the frequency of it s occurrence in a long (infinite) series of repeats P( A) = A Ω Ω A 0 P It s s represented by a number in the
Realizacja systemów wbudowanych (embeded systems) w strukturach PSoC (Programmable System on Chip)
 Realizacja systemów wbudowanych (embeded systems) w strukturach PSoC (Programmable System on Chip) Embeded systems Architektura układów PSoC (Cypress) Możliwości bloków cyfrowych i analogowych Narzędzia
Realizacja systemów wbudowanych (embeded systems) w strukturach PSoC (Programmable System on Chip) Embeded systems Architektura układów PSoC (Cypress) Możliwości bloków cyfrowych i analogowych Narzędzia
LEARNING AGREEMENT FOR STUDIES
 LEARNING AGREEMENT FOR STUDIES The Student First and last name(s) Nationality E-mail Academic year 2014/2015 Study period 1 st semester 2 nd semester Study cycle Bachelor Master Doctoral Subject area,
LEARNING AGREEMENT FOR STUDIES The Student First and last name(s) Nationality E-mail Academic year 2014/2015 Study period 1 st semester 2 nd semester Study cycle Bachelor Master Doctoral Subject area,
 !850016! www.irs.gov/form8879eo. e-file www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C,
!850016! www.irs.gov/form8879eo. e-file www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C,
EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO
 ARKUSZ ZAWIERA INFORMACJE PRAWNIE CHRONIONE DO MOMENTU ROZPOCZĘCIA EGZAMINU! Miejsce na naklejkę dysleksja MJA-R1_1P-082 EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO MAJ ROK 2008 POZIOM ROZSZERZONY CZĘŚĆ I
ARKUSZ ZAWIERA INFORMACJE PRAWNIE CHRONIONE DO MOMENTU ROZPOCZĘCIA EGZAMINU! Miejsce na naklejkę dysleksja MJA-R1_1P-082 EGZAMIN MATURALNY Z JĘZYKA ANGIELSKIEGO MAJ ROK 2008 POZIOM ROZSZERZONY CZĘŚĆ I
archivist: Managing Data Analysis Results
 archivist: Managing Data Analysis Results https://github.com/pbiecek/archivist Marcin Kosiński 1,2, Przemysław Biecek 2 1 IT Research and Development Grupa Wirtualna Polska 2 Faculty of Mathematics, Informatics
archivist: Managing Data Analysis Results https://github.com/pbiecek/archivist Marcin Kosiński 1,2, Przemysław Biecek 2 1 IT Research and Development Grupa Wirtualna Polska 2 Faculty of Mathematics, Informatics
Ukryte funkcjonalności w oprogramowaniu i urządzeniach elektronicznych. mgr inż. Paweł Koszut
 Ukryte funkcjonalności w oprogramowaniu i urządzeniach elektronicznych mgr inż. Paweł Koszut Ukryte funkcjonalności w oprogramowaniu i urządzeniach elektronicznych Zamiast wstępu : Inspiracja GSM w Grecji
Ukryte funkcjonalności w oprogramowaniu i urządzeniach elektronicznych mgr inż. Paweł Koszut Ukryte funkcjonalności w oprogramowaniu i urządzeniach elektronicznych Zamiast wstępu : Inspiracja GSM w Grecji
Jazz EB207S is a slim, compact and outstanding looking SATA to USB 2.0 HDD enclosure. The case is
 1. Introduction Jazz EB207S is a slim, compact and outstanding looking SATA to USB 2.0 HDD enclosure. The case is made of aluminum and steel mesh as one of the coolest enclosures available. It s also small
1. Introduction Jazz EB207S is a slim, compact and outstanding looking SATA to USB 2.0 HDD enclosure. The case is made of aluminum and steel mesh as one of the coolest enclosures available. It s also small
The shape of and the challenges for the Polish EO sector initial findings of the SEED EO project
 The shape of and the challenges for the Polish EO sector initial findings of the SEED EO project Drugie Forum Obserwacji Ziemi Ministerstwo Rozwoju Warszawa, 4 lipca 2016 2 Zadania projektu Stworzenie
The shape of and the challenges for the Polish EO sector initial findings of the SEED EO project Drugie Forum Obserwacji Ziemi Ministerstwo Rozwoju Warszawa, 4 lipca 2016 2 Zadania projektu Stworzenie
Unit of Social Gerontology, Institute of Labour and Social Studies ageing and its consequences for society
 Prof. Piotr Bledowski, Ph.D. Institute of Social Economy, Warsaw School of Economics local policy, social security, labour market Unit of Social Gerontology, Institute of Labour and Social Studies ageing
Prof. Piotr Bledowski, Ph.D. Institute of Social Economy, Warsaw School of Economics local policy, social security, labour market Unit of Social Gerontology, Institute of Labour and Social Studies ageing
OPTYMALIZACJA PUBLICZNEGO TRANSPORTU ZBIOROWEGO W GMINIE ŚRODA WIELKOPOLSKA
 Politechnika Poznańska Wydział Maszyn Roboczych i Transportu Inż. NATALIA LEMTIS OPTYMALIZACJA PUBLICZNEGO TRANSPORTU ZBIOROWEGO W GMINIE ŚRODA WIELKOPOLSKA Promotor: DR INŻ. MARCIN KICIŃSKI Poznań, 2016
Politechnika Poznańska Wydział Maszyn Roboczych i Transportu Inż. NATALIA LEMTIS OPTYMALIZACJA PUBLICZNEGO TRANSPORTU ZBIOROWEGO W GMINIE ŚRODA WIELKOPOLSKA Promotor: DR INŻ. MARCIN KICIŃSKI Poznań, 2016
Wybrzeze Baltyku, mapa turystyczna 1: (Polish Edition)
 Wybrzeze Baltyku, mapa turystyczna 1:50 000 (Polish Edition) Click here if your download doesn"t start automatically Wybrzeze Baltyku, mapa turystyczna 1:50 000 (Polish Edition) Wybrzeze Baltyku, mapa
Wybrzeze Baltyku, mapa turystyczna 1:50 000 (Polish Edition) Click here if your download doesn"t start automatically Wybrzeze Baltyku, mapa turystyczna 1:50 000 (Polish Edition) Wybrzeze Baltyku, mapa
Poland) Wydawnictwo "Gea" (Warsaw. Click here if your download doesn"t start automatically
 Suwalski Park Krajobrazowy i okolice 1:50 000, mapa turystyczno-krajoznawcza =: Suwalki Landscape Park, tourist map = Suwalki Naturpark,... narodowe i krajobrazowe) (Polish Edition) Click here if your
Suwalski Park Krajobrazowy i okolice 1:50 000, mapa turystyczno-krajoznawcza =: Suwalki Landscape Park, tourist map = Suwalki Naturpark,... narodowe i krajobrazowe) (Polish Edition) Click here if your
Towards Stability Analysis of Data Transport Mechanisms: a Fluid Model and an Application
 Towards Stability Analysis of Data Transport Mechanisms: a Fluid Model and an Application Gayane Vardoyan *, C. V. Hollot, Don Towsley* * College of Information and Computer Sciences, Department of Electrical
Towards Stability Analysis of Data Transport Mechanisms: a Fluid Model and an Application Gayane Vardoyan *, C. V. Hollot, Don Towsley* * College of Information and Computer Sciences, Department of Electrical
INSTYTUT GENETYKI I HODOWLI ZWIERZĄT POLSKIEJ AKADEMII NAUK W JASTRZĘBCU. mgr inż. Ewa Metera-Zarzycka
 INSTYTUT GENETYKI I HODOWLI ZWIERZĄT POLSKIEJ AKADEMII NAUK W JASTRZĘBCU mgr inż. Ewa Metera-Zarzycka Profil metaboliczny osocza krwi i wartość biologiczna mleka krów w gospodarstwach ekologicznych Praca
INSTYTUT GENETYKI I HODOWLI ZWIERZĄT POLSKIEJ AKADEMII NAUK W JASTRZĘBCU mgr inż. Ewa Metera-Zarzycka Profil metaboliczny osocza krwi i wartość biologiczna mleka krów w gospodarstwach ekologicznych Praca
Leba, Rowy, Ustka, Slowinski Park Narodowy, plany miast, mapa turystyczna =: Tourist map = Touristenkarte (Polish Edition)
 Leba, Rowy, Ustka, Slowinski Park Narodowy, plany miast, mapa turystyczna =: Tourist map = Touristenkarte (Polish Edition) FotKart s.c Click here if your download doesn"t start automatically Leba, Rowy,
Leba, Rowy, Ustka, Slowinski Park Narodowy, plany miast, mapa turystyczna =: Tourist map = Touristenkarte (Polish Edition) FotKart s.c Click here if your download doesn"t start automatically Leba, Rowy,
DUAL SIMILARITY OF VOLTAGE TO CURRENT AND CURRENT TO VOLTAGE TRANSFER FUNCTION OF HYBRID ACTIVE TWO- PORTS WITH CONVERSION
 ELEKTRYKA 0 Zeszyt (9) Rok LX Andrzej KUKIEŁKA Politechnika Śląska w Gliwicach DUAL SIMILARITY OF VOLTAGE TO CURRENT AND CURRENT TO VOLTAGE TRANSFER FUNCTION OF HYBRID ACTIVE TWO- PORTS WITH CONVERSION
ELEKTRYKA 0 Zeszyt (9) Rok LX Andrzej KUKIEŁKA Politechnika Śląska w Gliwicach DUAL SIMILARITY OF VOLTAGE TO CURRENT AND CURRENT TO VOLTAGE TRANSFER FUNCTION OF HYBRID ACTIVE TWO- PORTS WITH CONVERSION
Installation of EuroCert software for qualified electronic signature
 Installation of EuroCert software for qualified electronic signature for Microsoft Windows systems Warsaw 28.08.2019 Content 1. Downloading and running the software for the e-signature... 3 a) Installer
Installation of EuroCert software for qualified electronic signature for Microsoft Windows systems Warsaw 28.08.2019 Content 1. Downloading and running the software for the e-signature... 3 a) Installer
Ocena zachowań prozdrowotnych w zakresie higieny jamy ustnej obywateli
 Uniwersytet Medyczny w Lublinie lek. dent. Adriana Kleinert Ocena zachowań prozdrowotnych w zakresie higieny jamy ustnej obywateli polskich przebywających za granicą Rozprawa na stopień doktora nauk medycznych
Uniwersytet Medyczny w Lublinie lek. dent. Adriana Kleinert Ocena zachowań prozdrowotnych w zakresie higieny jamy ustnej obywateli polskich przebywających za granicą Rozprawa na stopień doktora nauk medycznych
POLITYKA PRYWATNOŚCI / PRIVACY POLICY
 POLITYKA PRYWATNOŚCI / PRIVACY POLICY TeleTrade DJ International Consulting Ltd Sierpień 2013 2011-2014 TeleTrade-DJ International Consulting Ltd. 1 Polityka Prywatności Privacy Policy Niniejsza Polityka
POLITYKA PRYWATNOŚCI / PRIVACY POLICY TeleTrade DJ International Consulting Ltd Sierpień 2013 2011-2014 TeleTrade-DJ International Consulting Ltd. 1 Polityka Prywatności Privacy Policy Niniejsza Polityka
do wniosku o przyznanie hodowcy wyłącznego prawa do odmiany for Plant Breeder s Rights (PBR)
 KWESTIONARIUSZ do wniosku o przyznanie hodowcy wyłącznego prawa do odmiany for Plant Breeder s Rights (PBR) 1. Roślina (takson) Plant (taxon) 1.1. Polska nazwa Polish name 1.2. Łacińska nazwa Latin name
KWESTIONARIUSZ do wniosku o przyznanie hodowcy wyłącznego prawa do odmiany for Plant Breeder s Rights (PBR) 1. Roślina (takson) Plant (taxon) 1.1. Polska nazwa Polish name 1.2. Łacińska nazwa Latin name
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition)
 Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Wojewodztwo Koszalinskie: Obiekty i walory krajoznawcze (Inwentaryzacja krajoznawcza Polski) (Polish Edition) Robert Respondowski Click here if your download doesn"t start automatically Wojewodztwo Koszalinskie:
Raport bieżący: 44/2018 Data: g. 21:03 Skrócona nazwa emitenta: SERINUS ENERGY plc
 Raport bieżący: 44/2018 Data: 2018-05-23 g. 21:03 Skrócona nazwa emitenta: SERINUS ENERGY plc Temat: Zawiadomienie o zmianie udziału w ogólnej liczbie głosów w Serinus Energy plc Podstawa prawna: Inne
Raport bieżący: 44/2018 Data: 2018-05-23 g. 21:03 Skrócona nazwa emitenta: SERINUS ENERGY plc Temat: Zawiadomienie o zmianie udziału w ogólnej liczbie głosów w Serinus Energy plc Podstawa prawna: Inne
Tomato Powdery Mildew (Leveillula taurica, Oidiopsis sicula)
 Tomato Powdery Mildew (Leveillula taurica, Oidiopsis sicula) Photo by Scott Stoddard Yield impact? Drying out/loss of foliage Sunburn of fruit No yield loss documented in processing tomatoes Fruit quality?
Tomato Powdery Mildew (Leveillula taurica, Oidiopsis sicula) Photo by Scott Stoddard Yield impact? Drying out/loss of foliage Sunburn of fruit No yield loss documented in processing tomatoes Fruit quality?
Effective Governance of Education at the Local Level
 Effective Governance of Education at the Local Level Opening presentation at joint Polish Ministry OECD conference April 16, 2012, Warsaw Mirosław Sielatycki Ministry of National Education Doskonalenie
Effective Governance of Education at the Local Level Opening presentation at joint Polish Ministry OECD conference April 16, 2012, Warsaw Mirosław Sielatycki Ministry of National Education Doskonalenie
Angielski Biznes Ciekawie
 Angielski Biznes Ciekawie Conditional sentences (type 2) 1. Discuss these two types of mindsets. 2. Decide how each type would act. 3. How would you act? Czy nauka gramatyki języka angielskiego jest trudna?
Angielski Biznes Ciekawie Conditional sentences (type 2) 1. Discuss these two types of mindsets. 2. Decide how each type would act. 3. How would you act? Czy nauka gramatyki języka angielskiego jest trudna?
Has the heat wave frequency or intensity changed in Poland since 1950?
 Has the heat wave frequency or intensity changed in Poland since 1950? Joanna Wibig Department of Meteorology and Climatology, University of Lodz, Poland OUTLINE: Motivation Data Heat wave frequency measures
Has the heat wave frequency or intensity changed in Poland since 1950? Joanna Wibig Department of Meteorology and Climatology, University of Lodz, Poland OUTLINE: Motivation Data Heat wave frequency measures
Knovel Math: Jakość produktu
 Knovel Math: Jakość produktu Knovel jest agregatorem materiałów pełnotekstowych dostępnych w formacie PDF i interaktywnym. Narzędzia interaktywne Knovel nie są stworzone wokół specjalnych algorytmów wymagających
Knovel Math: Jakość produktu Knovel jest agregatorem materiałów pełnotekstowych dostępnych w formacie PDF i interaktywnym. Narzędzia interaktywne Knovel nie są stworzone wokół specjalnych algorytmów wymagających
ROZPRAWA DOKTORSKA. Mateusz Romanowski
 Mateusz Romanowski Wpływ krioterapii ogólnoustrojowej na aktywność choroby i sprawność chorych na zesztywniające zapalenie stawów kręgosłupa ROZPRAWA DOKTORSKA Promotor: Dr hab., prof. AWF Anna Straburzyńska-Lupa
Mateusz Romanowski Wpływ krioterapii ogólnoustrojowej na aktywność choroby i sprawność chorych na zesztywniające zapalenie stawów kręgosłupa ROZPRAWA DOKTORSKA Promotor: Dr hab., prof. AWF Anna Straburzyńska-Lupa
Convolution semigroups with linear Jacobi parameters
 Convolution semigroups with linear Jacobi parameters Michael Anshelevich; Wojciech Młotkowski Texas A&M University; University of Wrocław February 14, 2011 Jacobi parameters. µ = measure with finite moments,
Convolution semigroups with linear Jacobi parameters Michael Anshelevich; Wojciech Młotkowski Texas A&M University; University of Wrocław February 14, 2011 Jacobi parameters. µ = measure with finite moments,
Formularz recenzji magazynu. Journal of Corporate Responsibility and Leadership Review Form
 Formularz recenzji magazynu Review Form Identyfikator magazynu/ Journal identification number: Tytuł artykułu/ Paper title: Recenzent/ Reviewer: (imię i nazwisko, stopień naukowy/name and surname, academic
Formularz recenzji magazynu Review Form Identyfikator magazynu/ Journal identification number: Tytuł artykułu/ Paper title: Recenzent/ Reviewer: (imię i nazwisko, stopień naukowy/name and surname, academic
Mgr Paweł Musiał. Promotor Prof. dr hab. n. med. Hanna Misiołek Promotor pomocniczy Dr n. med. Marek Tombarkiewicz
 Mgr Paweł Musiał Porównanie funkcjonowania podstawow-ych i specjalistycznych Zespołów Ratownictwa Medycznego na przykładzie Samodzielnego Publicznego Zespołu Zakładów Opieki Zdrowotnej w Staszowie Rozprawa
Mgr Paweł Musiał Porównanie funkcjonowania podstawow-ych i specjalistycznych Zespołów Ratownictwa Medycznego na przykładzie Samodzielnego Publicznego Zespołu Zakładów Opieki Zdrowotnej w Staszowie Rozprawa
Instrukcja obsługi User s manual
 Instrukcja obsługi User s manual Konfigurator Lanberg Lanberg Configurator E-mail: support@lanberg.pl support@lanberg.eu www.lanberg.pl www.lanberg.eu Lanberg 2015-2018 WERSJA VERSION: 2018/11 Instrukcja
Instrukcja obsługi User s manual Konfigurator Lanberg Lanberg Configurator E-mail: support@lanberg.pl support@lanberg.eu www.lanberg.pl www.lanberg.eu Lanberg 2015-2018 WERSJA VERSION: 2018/11 Instrukcja
 www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
www.irs.gov/form990. If "Yes," complete Schedule A Schedule B, Schedule of Contributors If "Yes," complete Schedule C, Part I If "Yes," complete Schedule C, Part II If "Yes," complete Schedule C, Part
Dolny Slask 1: , mapa turystycznosamochodowa: Plan Wroclawia (Polish Edition)
 Dolny Slask 1:300 000, mapa turystycznosamochodowa: Plan Wroclawia (Polish Edition) Click here if your download doesn"t start automatically Dolny Slask 1:300 000, mapa turystyczno-samochodowa: Plan Wroclawia
Dolny Slask 1:300 000, mapa turystycznosamochodowa: Plan Wroclawia (Polish Edition) Click here if your download doesn"t start automatically Dolny Slask 1:300 000, mapa turystyczno-samochodowa: Plan Wroclawia
Zestawienie czasów angielskich
 Zestawienie czasów angielskich Present Continuous I am, You are, She/ He/ It is, We/ You/ They are podmiot + operator + (czasownik główny + ing) + reszta I' m driving. operator + podmiot + (czasownik główny
Zestawienie czasów angielskich Present Continuous I am, You are, She/ He/ It is, We/ You/ They are podmiot + operator + (czasownik główny + ing) + reszta I' m driving. operator + podmiot + (czasownik główny
Institutional Determinants of IncomeLevel Convergence in the European. Union: Are Institutions Responsible for Divergence Tendencies of Some
 Institutional Determinants of IncomeLevel Convergence in the European Union: Are Institutions Responsible for Divergence Tendencies of Some Countries? Dr Mariusz Próchniak Katedra Ekonomii II, Szkoła Główna
Institutional Determinants of IncomeLevel Convergence in the European Union: Are Institutions Responsible for Divergence Tendencies of Some Countries? Dr Mariusz Próchniak Katedra Ekonomii II, Szkoła Główna
Employment. Number of employees employed on a contract of employment by gender in 2012. Company
 Im not found /sites/eneacsr2012.mess-asp.com/themes/eneacsr2012/img/enea.jpg Employt Capital Group is one of the largest companies in the energy industry. Therefore it has an influence, as an employer,
Im not found /sites/eneacsr2012.mess-asp.com/themes/eneacsr2012/img/enea.jpg Employt Capital Group is one of the largest companies in the energy industry. Therefore it has an influence, as an employer,
